MEDINRIA VIDE REGISTRATION
Registration of ( a) MRI images from a grapevine cutting: structures present at day 0 (red), day 35 (green) and at both dates (yellow) ( b) X-ray (red) and MRI (green) images from a human abdomen ( c) photographs (left), X-rays (center) and MRI (right) images from a grapevine trunk, in transversal (up) and longitudinal (down) views ( d) 3D time series from a living grapevine cutting acquired by MRI, using a mask to prevent local tissue deformation from being corrected ( Supplementary Data).
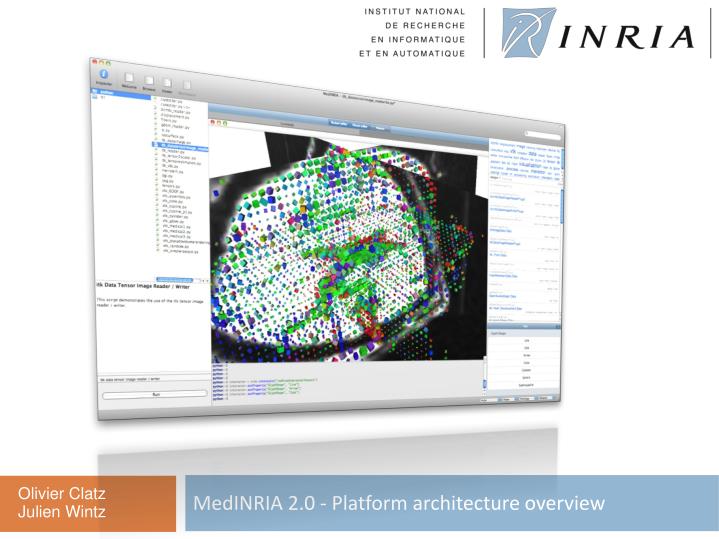
Validated transformations (matrices and/or deformation fields) and transformed images can be saved and reused either in the plugin or in Fiji. After registration, superimposition can be investigated visually, or quantitatively, using the mismatch measurement tool. Two optimization strategies can be applied: Block Matching or Iconic registration.
MEDINRIA VIDE MANUAL
Fijiyama proceeds with refining steps including an optional manual alignment in a tridimensional space for large deformations ( Fig. 1a), and automatic registration steps estimating rigid transformations, similarities or deformation fields ( Supplementary Data). Two pairwise image registration: Registration means estimating the geometrical transformation of a moving image to achieve the best possible superimposition with a reference image. Fijiyama addresses this concern, deriving biomedical algorithmic developments to build a generic registration plugin for Fiji, enabling multimodal and time-lapse 3D image registration by non-specialists. In ImageJ/Fiji ( Schindelin et al., 2012), existing plugins imply either human assistance for 3D registration (Atlas toolkit, Grocott et al., 2016), coding skills to achieve analysis of image series (Elastix, Klein et al., 2010), or are limited to handling a single imaging modality. While these two families of methods gave rise to several 3D registration tools, spanning from intermodal registration in human medicine ( Kikinis et al., 2014 Klein et al., 2010 Vichot et al., 2012 Yoo et al., 2002) to time-lapse tracking in plant imaging ( Fernandez et al., 2010 Michelin et al., 2016), no comprehensive and user-friendly solution is yet available to achieve both goals. Other registration strategies bypass modality-specific segmentation or points detection steps: (i) Iconic methods involve direct image comparisons computing pixel-based similarity metrics ( Beg et al., 2005 Vercauteren et al., 2009) (ii) Hybrid methods combine Iconic and segmentation-based strategies ( Commowick et al., 2006 Ourselin et al., 2000). However, this strategy may be of limited reliability when comparing dramatically different images collected from different modalities. Salient point detection methods were proposed in medical image registration ( Pennec et al., 2009), and used in microscopy image reconstruction ( Hörl et al., 2019).
This method is well suited for mono-modal reconstruction, but fiducial markers are modality-dependent. Correspondence points can be detected automatically using fiducial markers ( Preibisch et al., 2010), and paired using local descriptors. But incorrect point positioning leads to alignment mismatches, and the 3D segmentation task is sometimes time-consuming. Registration of 3D images can be performed by identifying correspondences between points ( Peng et al., 2011) or segmentations ( Grocott et al., 2016) defined by the user. Its versatility was assessed on four case studies combining multimodal and time series data, spanning from micro to macro scales. Fijiyama, a Fiji plugin built upon biomedical registration algorithms, is aimed at non-specialists to facilitate automatic alignment of 3D images acquired either at successive times and/or with different imaging systems. Identifying image invariants over modalities is challenging and can result in intractable problems. This registration step becomes more complex when combining observations from devices that highlight various tissue structures. Manual positioning and natural growth of the living samples induce variations in the shape, position and orientation in the acquired images that require a preprocessing step of 3D registration prior to analyses. However, living samples cannot remain in these devices for a long period. The anatomy, structure and function of tissues can be observed non-destructively in time-lapse multimodal imaging experiments by combining the outputs of imaging devices such as X-ray CT and MRI scans.
The increasing interest of animal and plant research communities for biomedical 3D imaging devices results in the emergence of new topics.
